Paired Model — Pólya-Gamma Gibbs Sampler¶
Overview¶
This model uses the same paired logistic regression as the Laplace variant, but performs exact posterior inference via Pólya-Gamma data augmentation and Gibbs sampling. It provides multi-chain MCMC diagnostics (R-hat, ESS) to verify convergence.
Generative model¶
Directed Acyclic Graph (DAG)¶
graph TD
sigma_mu(["σ_μ"]) --> mu["μ"]
sigma_delta(["σ_δ"]) --> delta_A["δ_A"]
mu --> pA["p_A = σ(μ + δ_A)"]
delta_A --> pA
mu --> pB["p_B = σ(μ)"]
pA --> yA(["y_A,i"])
pB --> yB(["y_B,i"])
style sigma_mu fill:#e0e0e0,stroke:#757575
style sigma_delta fill:#e0e0e0,stroke:#757575
style mu fill:#bbdefb,stroke:#1565c0
style delta_A fill:#bbdefb,stroke:#1565c0
style pA fill:#c8e6c9,stroke:#2e7d32
style pB fill:#c8e6c9,stroke:#2e7d32
style yA fill:#fff9c4,stroke:#f9a825
style yB fill:#fff9c4,stroke:#f9a825Legend: grey = hyperparameters, blue = latent parameters, green = deterministic, yellow = observed data.
Pólya-Gamma augmentation¶
Background¶
A random variable \(\omega \sim \text{PG}(b, c)\) follows a Pólya-Gamma distribution with parameters \(b > 0\) and \(c \in \mathbb{R}\). Its key property (Polson, Scott & Windle, 2013) is an integral identity that turns the logistic likelihood into a Gaussian scale mixture:
For a single Bernoulli observation \(y_i \in \{0,1\}\) with \(p_i = \sigma(\psi_i) = (1+e^{-\psi_i})^{-1}\), we have \(b=1\) and \(a = y_i\), giving \(\kappa_i = y_i - \tfrac{1}{2}\).
Likelihood rewrite¶
The full Bernoulli log-likelihood can be rewritten as:
Applying the PG integral identity to each factor \(\cosh(\psi_i/2)^{-1}\) introduces latent variables \(\omega_i \sim \text{PG}(1, 0)\) and yields a conditionally Gaussian augmented likelihood:
where \(\boldsymbol{\Omega} = \operatorname{diag}(\omega_1, \dots, \omega_{2n})\). Combined with the Gaussian prior on \(\boldsymbol{\beta}\), this gives closed-form full conditionals for both \(\boldsymbol{\beta}\) and \(\boldsymbol{\omega}\), which is the basis of the Gibbs sampler below.
Stacked design¶
We stack the \(n\) paired observations into a \(2n\)-dimensional regression:
so the linear predictor is \(\boldsymbol{\psi} = \mathbf{X}\boldsymbol{\beta}\), i.e. \(\psi_i = \mu + \delta_A\) for the group-A rows and \(\psi_i = \mu\) for the group-B rows, and \(\boldsymbol{\kappa} = \mathbf{y} - \tfrac{1}{2}\).
Prior¶
The Gaussian prior on \(\boldsymbol{\beta}\) is:
with prior precision \(\mathbf{B}_0^{-1} = \operatorname{diag}\!\bigl(1/\sigma_\mu^{2},\;1/\sigma_\delta^{2}\bigr)\).
Gibbs sampler¶
After augmenting with \(\omega_i \sim \text{PG}(1, \psi_i)\), the joint posterior of \((\boldsymbol{\beta}, \boldsymbol{\omega})\) admits two tractable full conditionals that are alternated at each sweep:
Step 1 — Sample auxiliary variables:
Step 2 — Sample regression coefficients:
where the posterior precision and mean are:
This is a standard Bayesian weighted least-squares update: the PG variables \(\omega_i\) act as observation weights. Because both conditionals are available in closed form, the sampler requires no tuning parameters (unlike Metropolis-Hastings) and mixes well for logistic models.
Why it works¶
Marginalising over \(\boldsymbol{\omega}\) recovers the exact logistic likelihood, so the marginal posterior \(p(\boldsymbol{\beta} \mid \mathbf{y})\) from the Gibbs sampler targets the true posterior — there is no approximation error beyond finite MCMC variance.
When to use¶
- Exact inference — no approximation error, exact up to MCMC error
- Convergence diagnostics — R-hat and ESS across multiple chains
- Final analysis — when you need trustworthy results for reporting
Note
The PG sampler is slower than the Laplace approximation. For exploration,
start with PairedBayesPropTest and switch to PairedBayesPropTestPG
for final analysis.
Step-by-step example¶
1. Simulate paired data¶
import numpy as np
from bayesprop.resources.bayes_paired_pg import PairedBayesPropTestPG, sigmoid
from bayesprop.resources.bayes_paired_laplace import PairedBayesPropTest
from bayesprop.utils.utils import simulate_paired_scores
sim = simulate_paired_scores(N=250, delta_A=1.0, sigma_theta=0.0, seed=42)
y_A = sim.y_A
y_B = sim.y_B
print(f"True δ_A = {sim.true_params.delta_A}")
print(f"Fraction y=1: A={y_A.mean():.1%}, B={y_B.mean():.1%}")
2. Fit the PG Gibbs model¶
pg_model = PairedBayesPropTestPG(
prior_sigma_delta=1.0,
prior_sigma_mu=2.0,
seed=42,
n_iter=2000,
burn_in=500,
n_chains=4,
).fit(y_A, y_B)
s = pg_model.summary
print(f"δ_A posterior mean = {s.delta_A_posterior_mean:+.4f}")
print(f"Mean Δ (prob) = {s.mean_delta:+.4f}")
print(f"95% CI = [{s.ci_95.lower:.4f}, {s.ci_95.upper:.4f}]")
print(f"P(A>B) = {s.p_A_greater_B:.4f}")
3. Unified decision¶
d = pg_model.decide()
print(f"Bayes Factor: BF₁₀ = {d.bayes_factor.BF_10:.2f} → {d.bayes_factor.decision}")
print(f"Posterior Null: P(H₀|D) = {d.posterior_null.p_H0:.4f} → {d.posterior_null.decision}")
print(f"ROPE: {d.rope.decision} ({d.rope.pct_in_rope:.1%} in ROPE)")
Bayes Factor: BF₁₀ = 10^27 → Reject H0
Posterior Null: P(H₀|D) = 0.0000 → Reject H0
ROPE: Reject H0 — A practically better (0.0% in ROPE)
4. MCMC diagnostics¶
Two standard diagnostics summarise sampler health:
- R-hat (Gelman-Rubin) compares between- and within-chain variance; values close to 1 (target \(\hat{R} < 1.05\)) indicate the chains have mixed to the same distribution.
- ESS (effective sample size) is the number of independent draws equivalent to the autocorrelated MCMC sample; target ESS > 400 per parameter for stable posterior summaries.
Convergence checks
Always verify that R-hat < 1.05 and ESS > 400 before
trusting the results. If convergence is poor, increase n_iter
or n_chains.
diag = pg_model.mcmc_diagnostics()
print(f"μ: R-hat={diag.mu.r_hat:.3f}, ESS={diag.mu.ess:.0f}")
print(f"δ_A: R-hat={diag.delta_A.r_hat:.3f}, ESS={diag.delta_A.ess:.0f}")
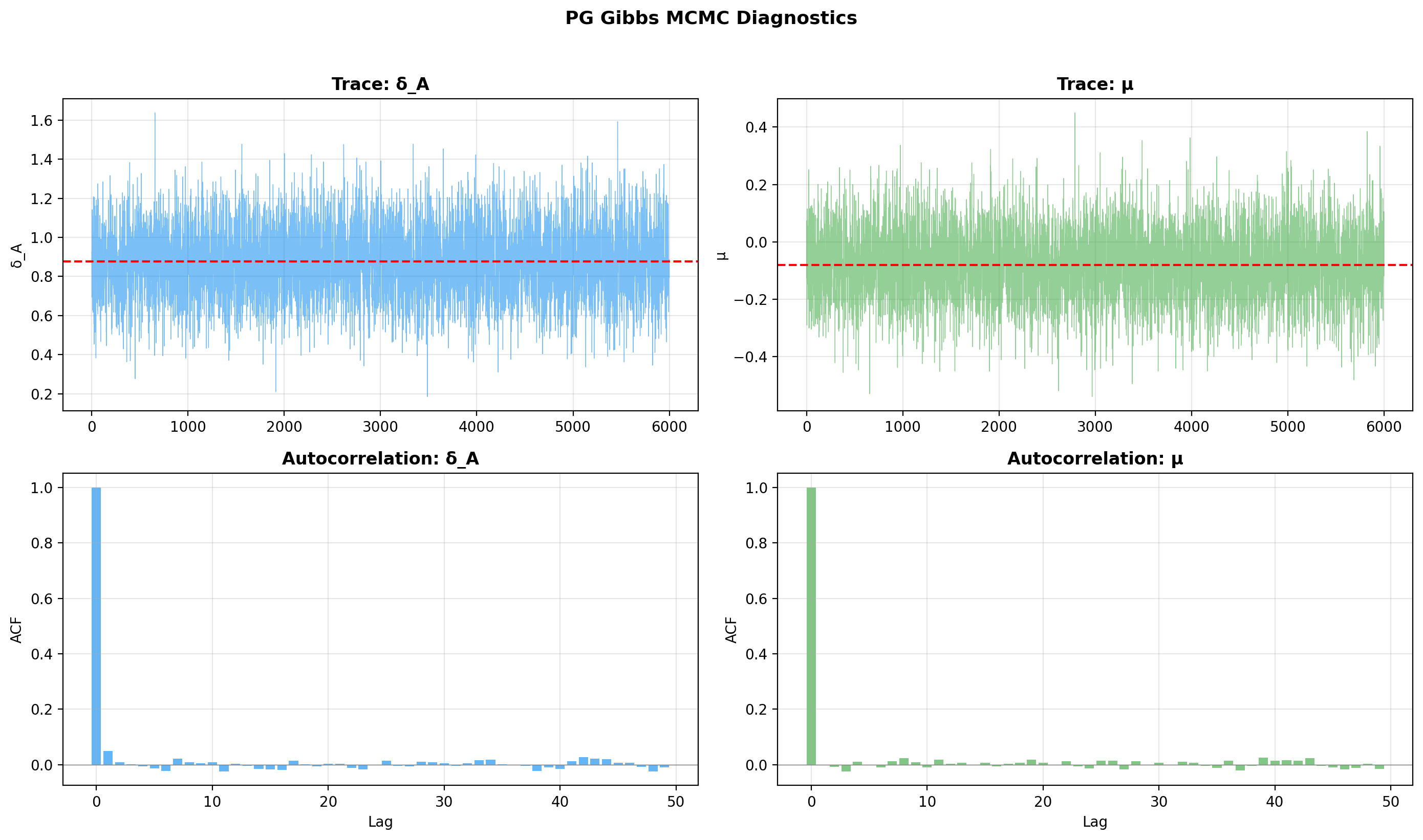
5. Trace and autocorrelation plots¶
Use the built-in trace plot — it renders all chains together, with per-parameter trace and ACF panels:
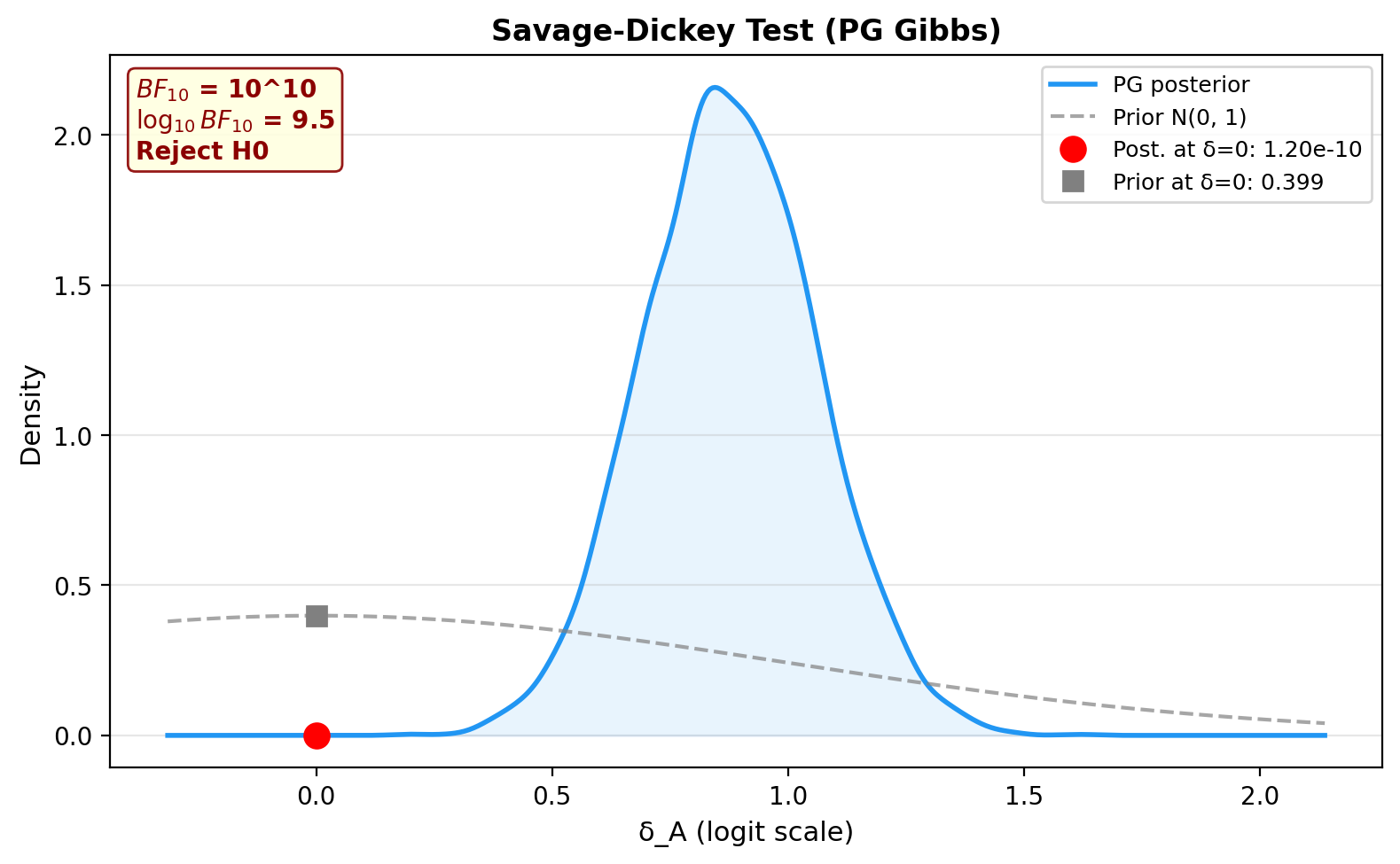
6. Savage-Dickey Bayes Factor¶
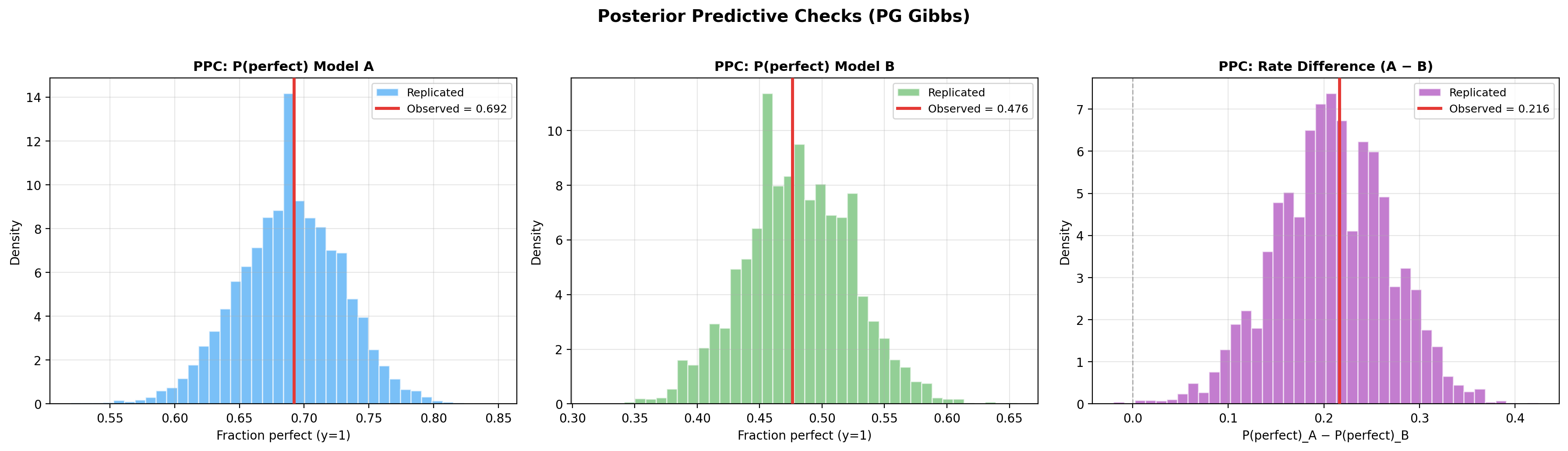
7. Posterior predictive checks¶
pg_model.plot_ppc(seed=42)
ppc = pg_model.ppc_pvalues(seed=42)
print(f"{'Statistic':<20} {'Observed':>10} {'p-value':>10} {'Status':>10}")
print("-" * 55)
for stat_name, vals in ppc.items():
print(f"{stat_name:<20} {vals.observed:>10.4f} {vals.p_value:>10.3f} {vals.status:>10}")
Comparison with Laplace¶
A key use case for the PG sampler is to verify that the Laplace approximation gives similar results. Fit both on the same data and compare:
laplace_model = PairedBayesPropTest(seed=42, n_samples=2000).fit(y_A, y_B)
delta_samples = pg_model.delta_A_samples
mu_samples = pg_model.samples[:, 0]
laplace_delta = laplace_model.delta_A_samples
laplace_mu = laplace_model.laplace["mu_samples"]
print("PG Gibbs vs Laplace — posterior summary")
print("=" * 55)
print(f"{'':20} {'PG Gibbs':>15} {'Laplace':>15}")
print("-" * 55)
print(f"{'δ_A mean':20} {delta_samples.mean():>15.4f} {laplace_delta.mean():>15.4f}")
print(f"{'δ_A sd':20} {delta_samples.std():>15.4f} {laplace_delta.std():>15.4f}")
print(f"{'μ mean':20} {mu_samples.mean():>15.4f} {laplace_mu.mean():>15.4f}")
print("=" * 55)
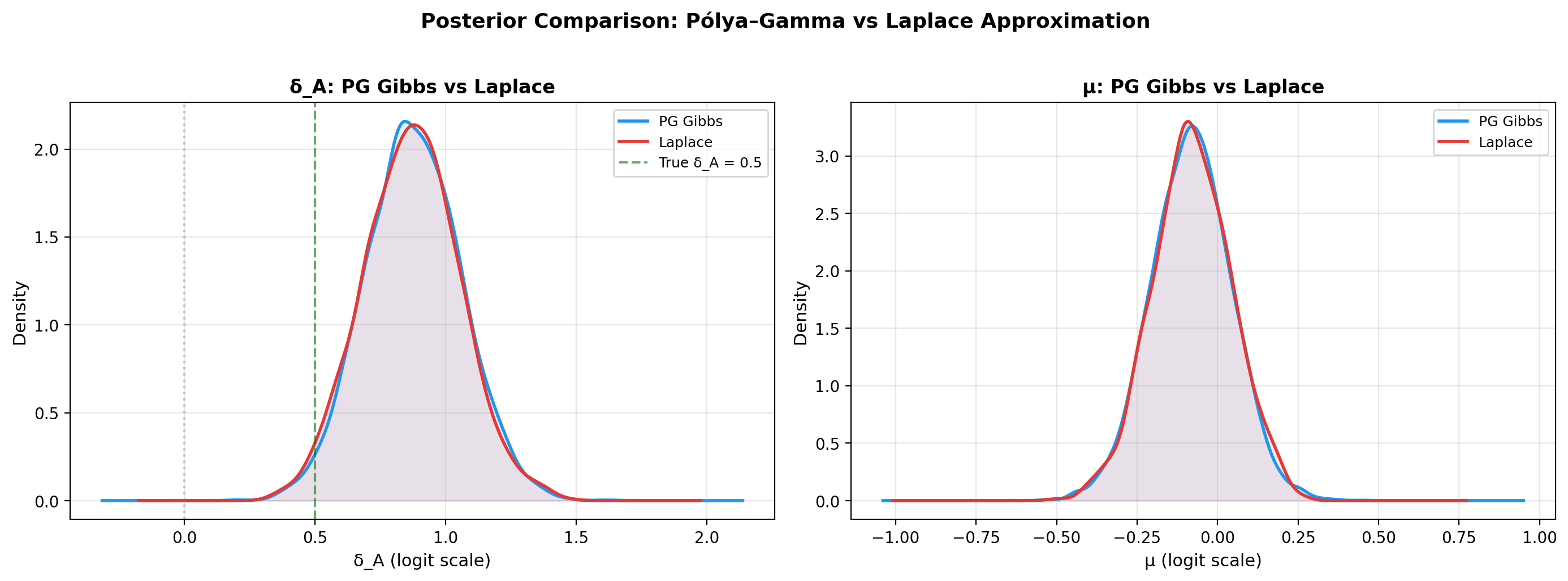
PG Gibbs vs Laplace: posterior KDEs of μ and δ_A¶
import matplotlib.pyplot as plt
from scipy.stats import gaussian_kde
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
# δ_A comparison
ax = axes[0]
for samples, label, color in [
(delta_samples, "PG Gibbs", "#2196F3"),
(laplace_delta, "Laplace", "#E53935"),
]:
kde = gaussian_kde(samples)
x = np.linspace(samples.min() - 0.5, samples.max() + 0.5, 300)
ax.plot(x, kde(x), linewidth=2, color=color, label=label)
ax.fill_between(x, kde(x), alpha=0.1, color=color)
ax.axvline(1.0, color="green", ls="--", alpha=0.6, label="True δ_A = 1.0")
ax.axvline(0, color="gray", ls=":", alpha=0.4)
ax.set_xlabel("δ_A (logit scale)")
ax.set_title("δ_A: PG Gibbs vs Laplace", fontweight="bold")
ax.legend(fontsize=9)
ax.grid(alpha=0.3)
# μ comparison
ax = axes[1]
for samples, label, color in [
(mu_samples, "PG Gibbs", "#2196F3"),
(laplace_mu, "Laplace", "#E53935"),
]:
kde = gaussian_kde(samples)
x = np.linspace(samples.min() - 0.5, samples.max() + 0.5, 300)
ax.plot(x, kde(x), linewidth=2, color=color, label=label)
ax.fill_between(x, kde(x), alpha=0.1, color=color)
ax.set_xlabel("μ (logit scale)")
ax.set_title("μ: PG Gibbs vs Laplace", fontweight="bold")
ax.legend(fontsize=9)
ax.grid(alpha=0.3)
fig.suptitle("Posterior Comparison: Pólya–Gamma vs Laplace",
fontsize=13, fontweight="bold", y=1.02)
plt.tight_layout()
plt.show()
Savage-Dickey comparison¶
d_pg = pg_model.decide()
d_lp = laplace_model.decide()
print(f"{'':20} {'PG Gibbs':>15} {'Laplace':>15}")
print("-" * 55)
print(f"{'BF₁₀':20} {d_pg.bayes_factor.BF_10:>15.2f} {d_lp.bayes_factor.BF_10:>15.2f}")
print(f"{'BF Decision':20} {d_pg.bayes_factor.decision:>15} {d_lp.bayes_factor.decision:>15}")
print(f"{'ROPE Decision':20} {d_pg.rope.decision:>15} {d_lp.rope.decision:>15}")
| Aspect | Laplace | Pólya-Gamma |
|---|---|---|
| Speed | Fast (milliseconds) | Slower (seconds) |
| Accuracy | Approximate | Exact (up to MCMC) |
| Diagnostics | None | R-hat, ESS |
| Recommended for | Exploration | Final reporting |
Unified decision comparison¶
print("Unified Decision — PG Gibbs vs Laplace")
print("=" * 70)
for label, m in [("PG Gibbs", pg_model), ("Laplace", laplace_model)]:
d = m.decide()
bf = d.bayes_factor
pn = d.posterior_null
rp = d.rope
print(f"\n{label}:")
print(f" Bayes Factor: BF₁₀ = {bf.BF_10:.2f} → {bf.decision}")
print(f" Posterior Null: P(H₀|D) = {pn.p_H0:.4f} → {pn.decision}")
print(f" ROPE [{rp.rope_lower:.2f}, {rp.rope_upper:.2f}]: "
f"{rp.decision} ({rp.pct_in_rope:.1%} in ROPE)")
BFDA sample-size planning¶
from bayesprop.utils.utils import (
bfda_power_curve,
plot_bfda_power,
plot_bfda_sensitivity,
)
theta_A_hat = y_A.mean()
theta_B_hat = y_B.mean()
sample_sizes = [20, 30, 50, 75, 100, 150, 200, 300, 500]
power_curve = bfda_power_curve(
theta_A_true=theta_A_hat,
theta_B_true=theta_B_hat,
sample_sizes=sample_sizes,
design="paired",
decision_rule="bayes_factor",
bf_threshold=3.0,
n_sim=200,
n_iter=1000,
burn_in=300,
n_chains=2,
seed=42,
)
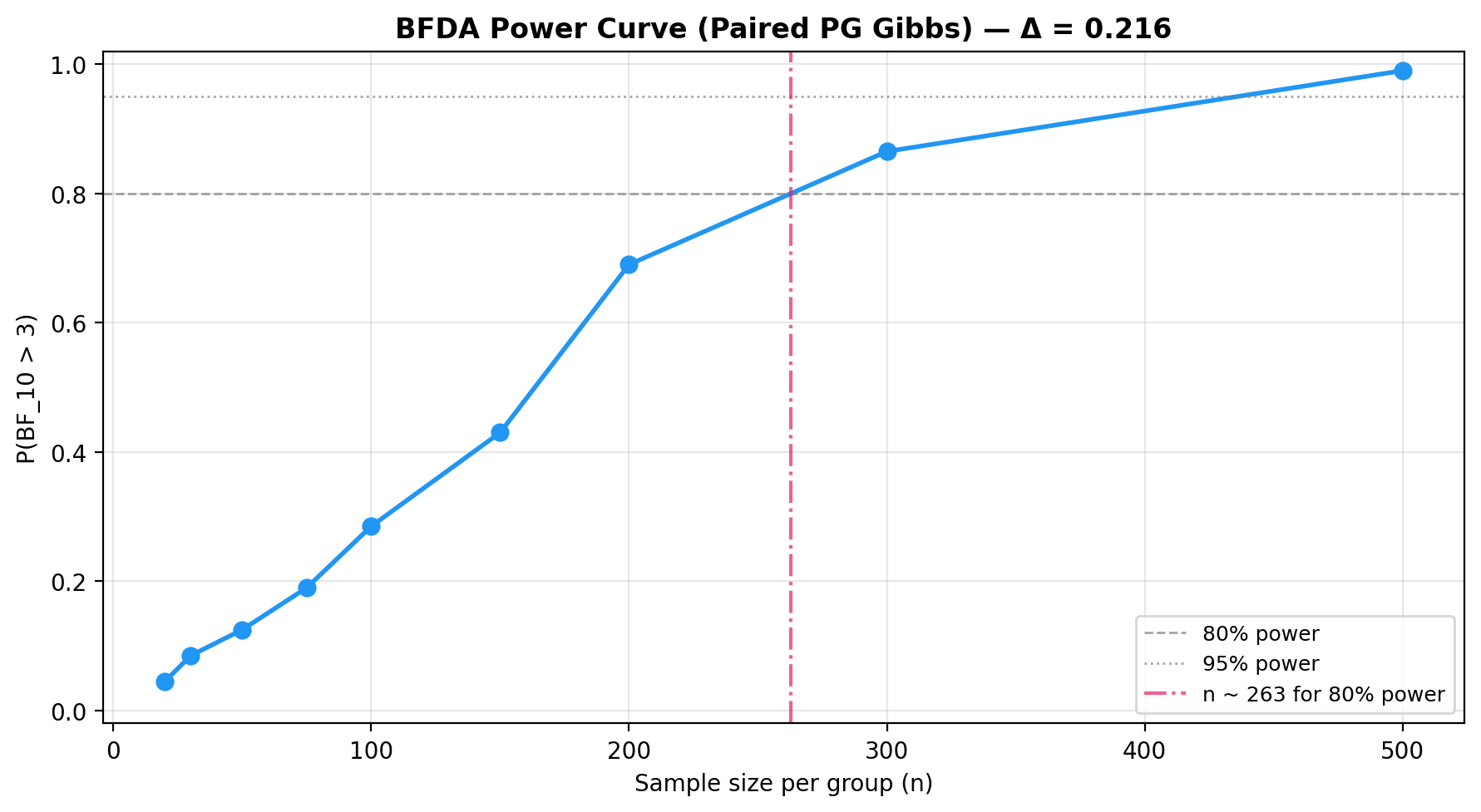
plot_bfda_power(
power_curve, theta_A_hat, theta_B_hat,
title=f"BFDA Power Curve (Paired PG Gibbs) — Δ = {theta_A_hat - theta_B_hat:.3f}"
)
Sensitivity to BF threshold:
plot_bfda_sensitivity(
theta_A_true=theta_A_hat,
theta_B_true=theta_B_hat,
sample_sizes=sample_sizes,
thresholds=[3.0, 6.0, 10.0],
n_sim=200,
seed=42,
design="paired",
title="BFDA Sensitivity — Multiple BF₁₀ Thresholds",
)
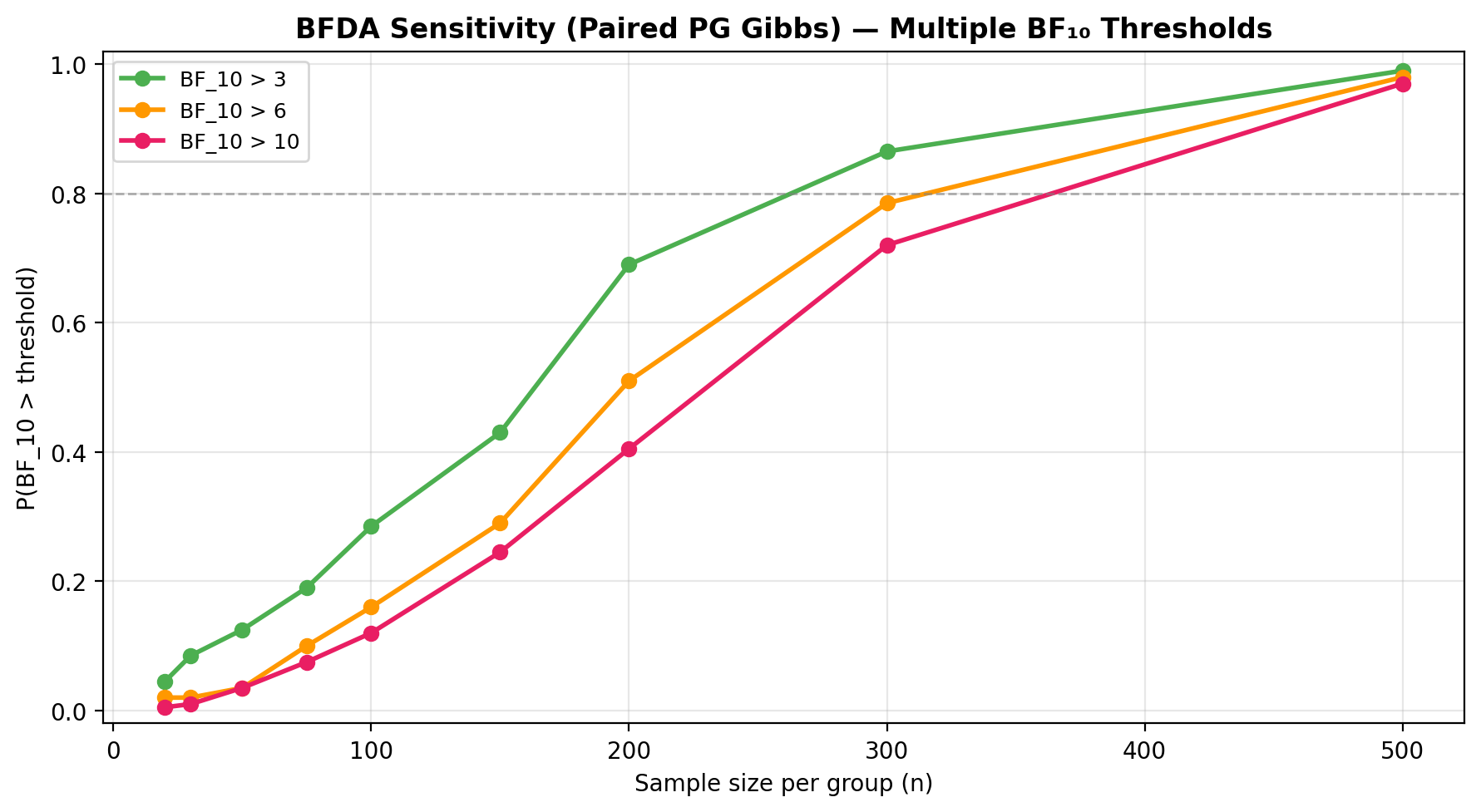
See the BFDA guide for the full sample-size planning workflow.
API¶
See API Reference — Paired Model (Pólya-Gamma) for full method documentation.
References¶
- Polson, N. G., Scott, J. G. & Windle, J. (2013). Bayesian inference for logistic models using Pólya-Gamma latent variables. Journal of the American Statistical Association, 108(504), 1339–1349.
- Windle, J., Polson, N. G. & Scott, J. G. (2014). Sampling Pólya-Gamma random variates: alternate and approximate techniques. arXiv:1405.0506.