Paired Model — Laplace Approximation¶
Overview¶
The paired model is used when both conditions are evaluated on the same items or subjects. It uses a pooled Bernoulli logistic regression with a Laplace approximation (MAP + analytical Hessian) for fast, analytic posterior inference.
Generative model¶
where \(\sigma(x) = 1/(1 + e^{-x})\) is the logistic sigmoid function. The parameter \(\delta_A\) captures group A's advantage on the logit scale; \(\mu\) is the shared baseline log-odds.
Directed Acyclic Graph (DAG)¶
graph TD
sigma_mu(["σ_μ"]) --> mu["μ"]
sigma_delta(["σ_δ"]) --> delta_A["δ_A"]
mu --> pA["p_A = σ(μ + δ_A)"]
delta_A --> pA
mu --> pB["p_B = σ(μ)"]
pA --> yA(["y_A,i"])
pB --> yB(["y_B,i"])
style sigma_mu fill:#e0e0e0,stroke:#757575
style sigma_delta fill:#e0e0e0,stroke:#757575
style mu fill:#bbdefb,stroke:#1565c0
style delta_A fill:#bbdefb,stroke:#1565c0
style pA fill:#c8e6c9,stroke:#2e7d32
style pB fill:#c8e6c9,stroke:#2e7d32
style yA fill:#fff9c4,stroke:#f9a825
style yB fill:#fff9c4,stroke:#f9a825Legend: grey = hyperparameters, blue = latent parameters, green = deterministic, yellow = observed data.
Laplace approximation¶
The Laplace method approximates the posterior as a bivariate Gaussian centred at the MAP (maximum a posteriori) estimate:
where \(\mathbf{H}\) is the Hessian of the negative log-posterior evaluated at the MAP.
Log-posterior¶
Let \(\mathbf{\theta} = (\mu, \delta_A)^\top\) denote the parameter vector. The log-posterior is
with \(p_A = \sigma(\mu + \delta_A)\) and \(p_B = \sigma(\mu)\).
Gradient¶
where \(k_A = \sum y_{A,i}\) and \(k_B = \sum y_{B,i}\).
Hessian of negative log-posterior¶
where \(w_A = p_A(1 - p_A)\) and \(w_B = p_B(1 - p_B)\), evaluated at the MAP.
Solver¶
The MAP is found by damped Newton iteration in 2D using the closed-form gradient and Hessian above (no external optimizer is invoked). Each step solves the \(2\times 2\) system \(\mathbf{H}\,\Delta\boldsymbol{\theta} = -\nabla(-\log p)\) in closed form via the cofactor inverse, and an Armijo backtracking line search guarantees monotone descent even from a poor starting point.
Because the negative log-posterior is strictly convex (Gaussian priors plus
Bernoulli likelihood), Newton converges quadratically. The
:class:SequentialPairedBayesPropTest warm-starts each look from the
previous MAP, which typically requires only 1–3 iterations per update.
When to use¶
- Fast inference — no MCMC, results in milliseconds
- Moderate sample sizes — works well with \(n \geq 30\)
- Exploratory analysis — quick iteration before committing to full MCMC
For exact posterior inference with convergence diagnostics, see Paired Model (Pólya-Gamma).
Step-by-step example¶
1. Simulate paired data¶
from bayesprop.resources.bayes_paired_laplace import PairedBayesPropTest
from bayesprop.utils.utils import simulate_paired_scores
sim = simulate_paired_scores(N=250, delta_A=0.8, sigma_theta=0.0, seed=42)
print(f"True δ_A = {sim.true_params.delta_A}")
print(f"Observed rates: A = {sim.y_A.mean():.3f}, B = {sim.y_B.mean():.3f}")
2. Fit the model¶
model = PairedBayesPropTest(
prior_sigma_delta=1.0,
seed=42,
n_samples=50_000,
).fit(sim.y_A, sim.y_B)
s = model.summary
print(f"δ_A posterior mean = {s.delta_A_posterior_mean:+.4f}")
print(f"Mean Δ (prob) = {s.mean_delta:+.4f}")
print(f"95% CI = [{s.ci_95.lower:.4f}, {s.ci_95.upper:.4f}]")
print(f"P(A>B) = {s.p_A_greater_B:.4f}")
3. Unified decision¶
d = model.decide()
print(f"Bayes Factor: BF₁₀ = {d.bayes_factor.BF_10:.2f} → {d.bayes_factor.decision}")
print(f"Posterior Null: P(H₀|D) = {d.posterior_null.p_H0:.4f} → {d.posterior_null.decision}")
print(f"ROPE: {d.rope.decision} ({d.rope.pct_in_rope:.1%} in ROPE)")
4. Laplace posterior visualisation¶
The Laplace approximation produces a bivariate Gaussian in \((\mu, \delta_A)\). Use the built-in method to inspect the implied probability posteriors \(p_A = \sigma(\mu + \delta_A)\), \(p_B = \sigma(\mu)\) and their difference \(\Delta = p_A - p_B\):
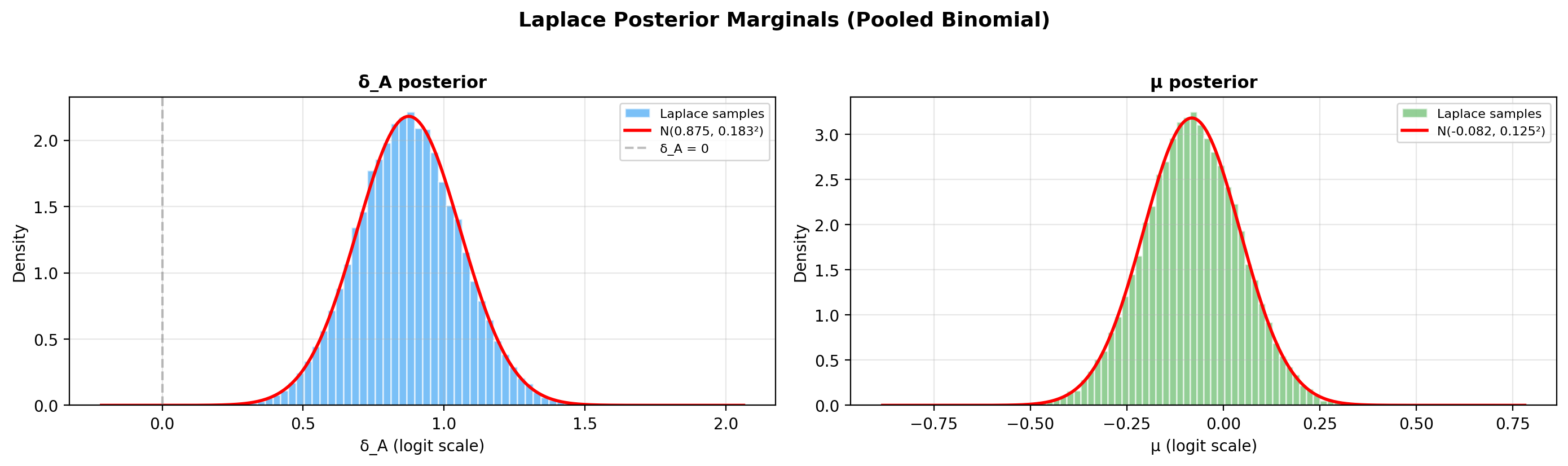
If you need the raw MAP / covariance values for a custom plot, they are available on the fitted model:
import numpy as np
laplace = model.laplace
mu_map, delta_map = laplace["map"]
cov = laplace["cov"]
sd_m, sd_d = np.sqrt(cov[0, 0]), np.sqrt(cov[1, 1])
print(f"MAP: μ={mu_map:.4f}, δ_A={delta_map:.4f}")
print(f"Posterior sd: μ={sd_m:.4f}, δ_A={sd_d:.4f}")
print(f"Correlation: {cov[0, 1] / (sd_m * sd_d):.3f}")
5. Posterior of Delta on the probability scale¶
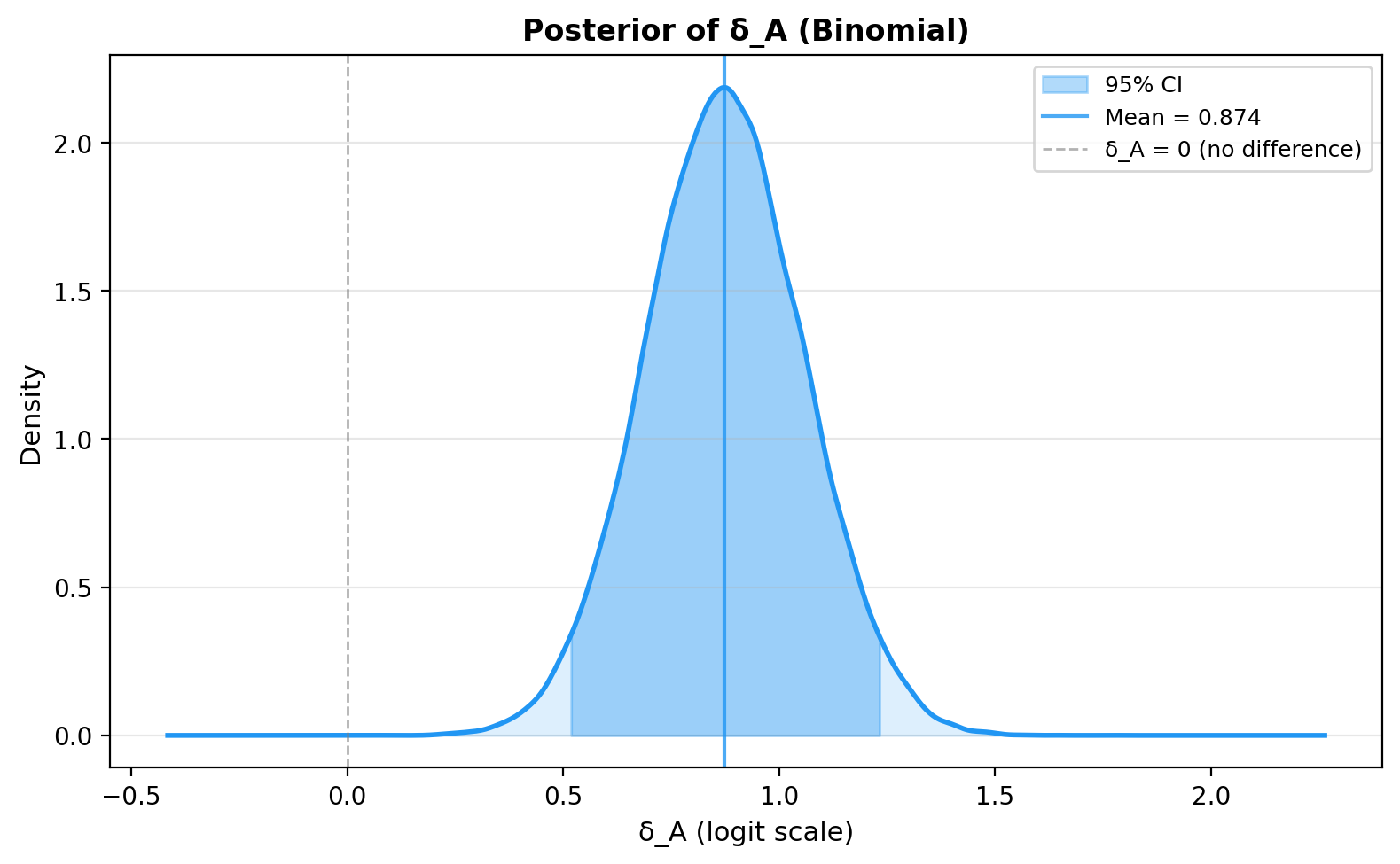
6. Savage-Dickey Bayes Factor plot¶
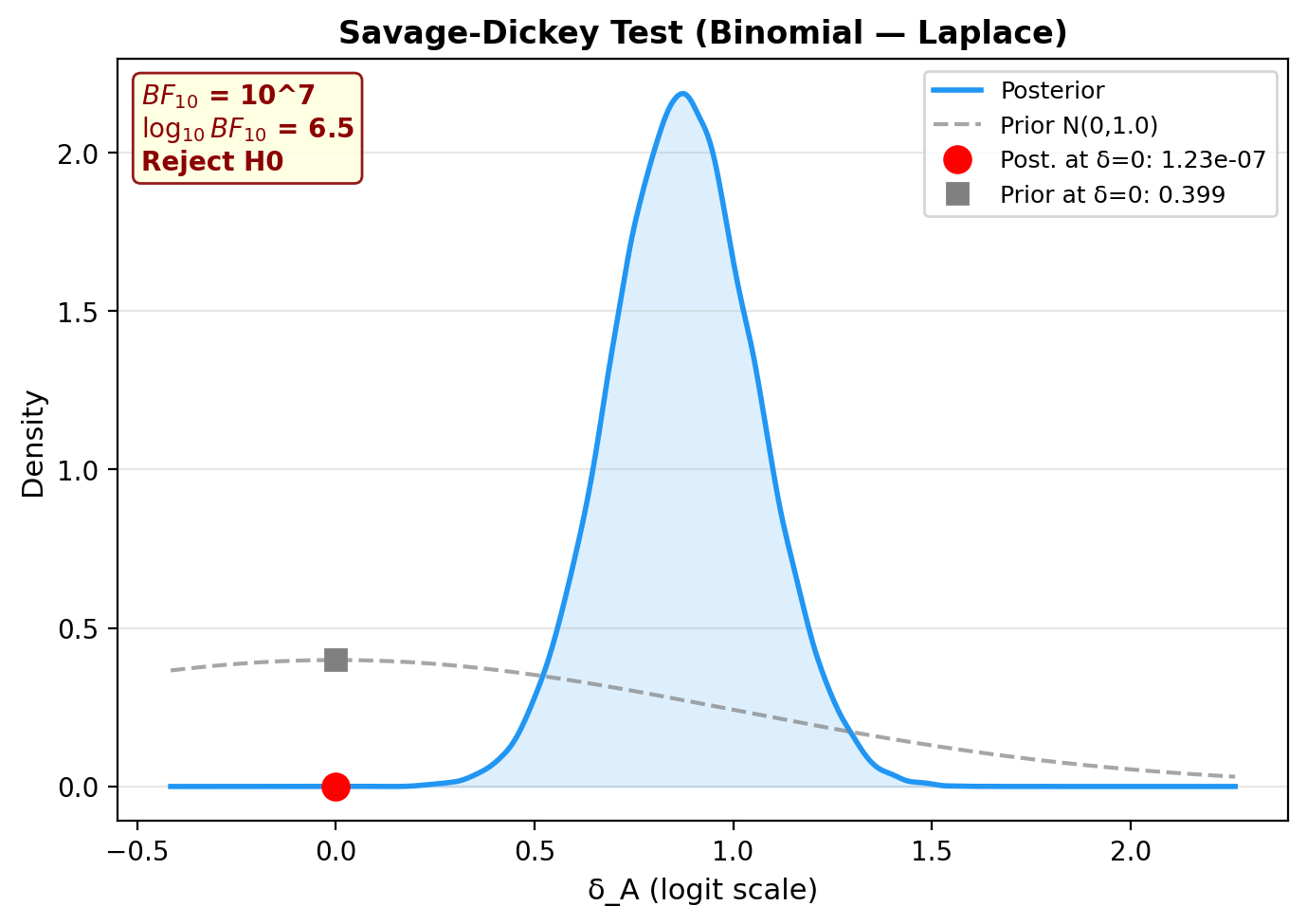
7. Posterior predictive checks¶
ppc = model.ppc_pvalues(seed=42)
print(f"{'Statistic':<20} {'Observed':>10} {'p-value':>10} {'Status':>10}")
print("-" * 55)
for stat_name, vals in ppc.items():
print(f"{stat_name:<20} {vals.observed:>10.4f} {vals.p_value:>10.3f} {vals.status:>10}")
PPC plots (fraction perfect for each model + rate difference):
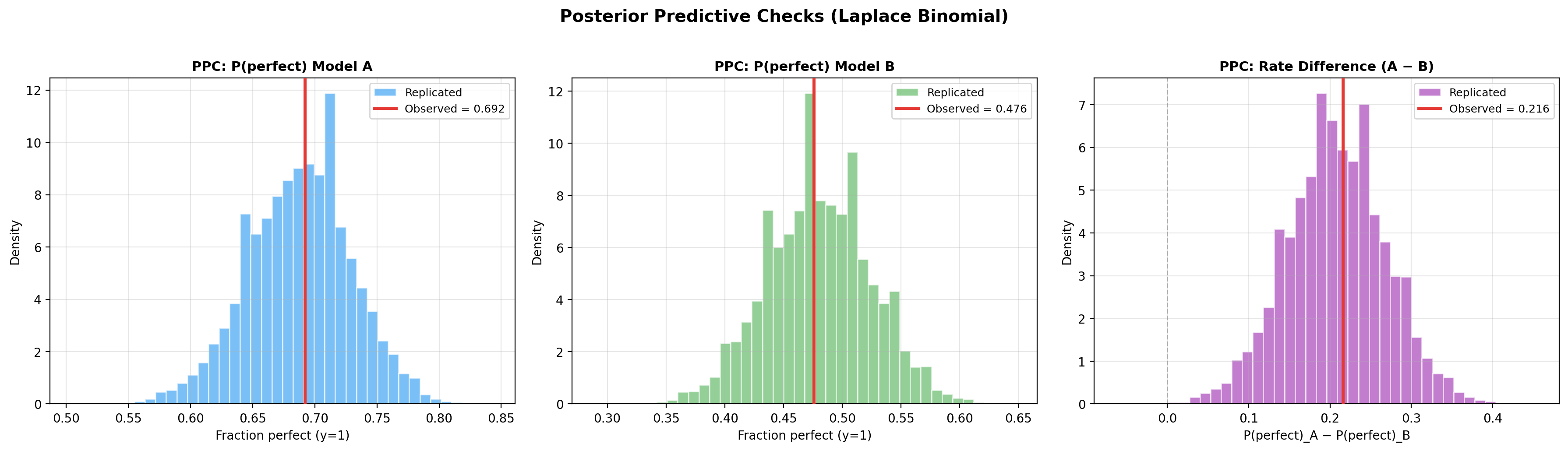
Prior sensitivity analysis¶
Sensitivity to prior P(H0)¶
Plot how the posterior \(P(H_0 \mid D)\) changes as you vary the prior \(\pi_0 = P(H_0)\):
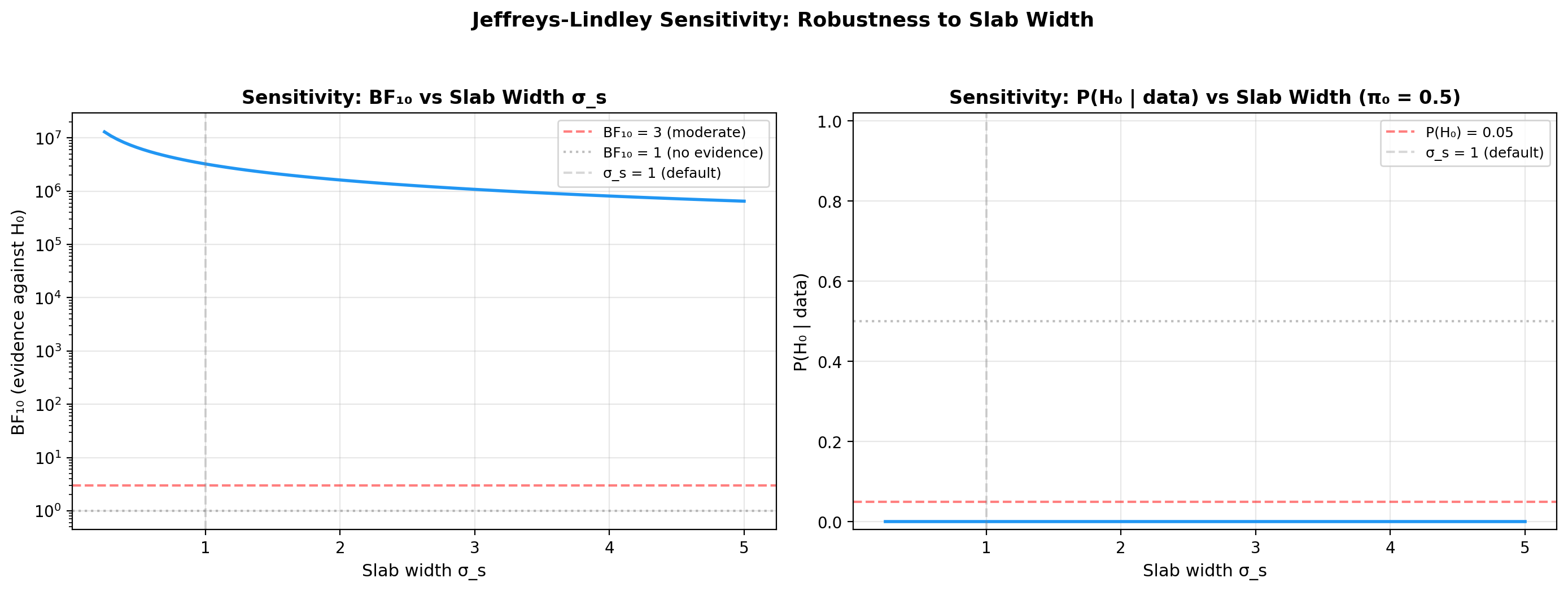
Sensitivity to slab width sigma_s¶
The Savage-Dickey BF depends on the prior at \(\delta_A = 0\). For a
\(\mathcal{N}(0, \sigma_s)\) slab prior, a wider slab concentrates less
density at zero, inflating \(BF_{10}\). This is the Jeffreys-Lindley
paradox in action. The right panel of plot_sensitivity above already
sweeps \(\sigma_s\) on a log scale, so no extra code is needed:
Frequentist comparison (McNemar test)¶
For reference, you can compare the Bayesian result with McNemar's test on the same binarized paired data:
from scipy.stats import chi2
import math
y_A = model.y_A_obs
y_B = model.y_B_obs
b = np.sum((y_A == 1) & (y_B == 0)) # A perfect, B not
c = np.sum((y_A == 0) & (y_B == 1)) # B perfect, A not
if b + c > 0:
chi2_stat = (b - c) ** 2 / (b + c)
p_val = 1 - chi2.cdf(chi2_stat, df=1)
print(f"Discordant pairs: A>B={b}, B>A={c}")
print(f"McNemar χ² = {chi2_stat:.2f}, p = {p_val:.4f}")
BFDA sample-size planning¶
from bayesprop.utils.utils import bfda_power_curve, plot_bfda_power
theta_A_hat = model.y_A_obs.mean()
theta_B_hat = model.y_B_obs.mean()
sample_sizes = [20, 30, 50, 75, 100, 150, 200, 300, 500]
power_curve = bfda_power_curve(
theta_A_true=theta_A_hat,
theta_B_true=theta_B_hat,
sample_sizes=sample_sizes,
design="paired",
decision_rule="bayes_factor",
bf_threshold=3.0,
n_sim=200,
seed=42,
)
plot_bfda_power(
power_curve, theta_A_hat, theta_B_hat,
title=f"BFDA Power Curve (Paired Laplace) — Δ = {theta_A_hat - theta_B_hat:.3f}"
)
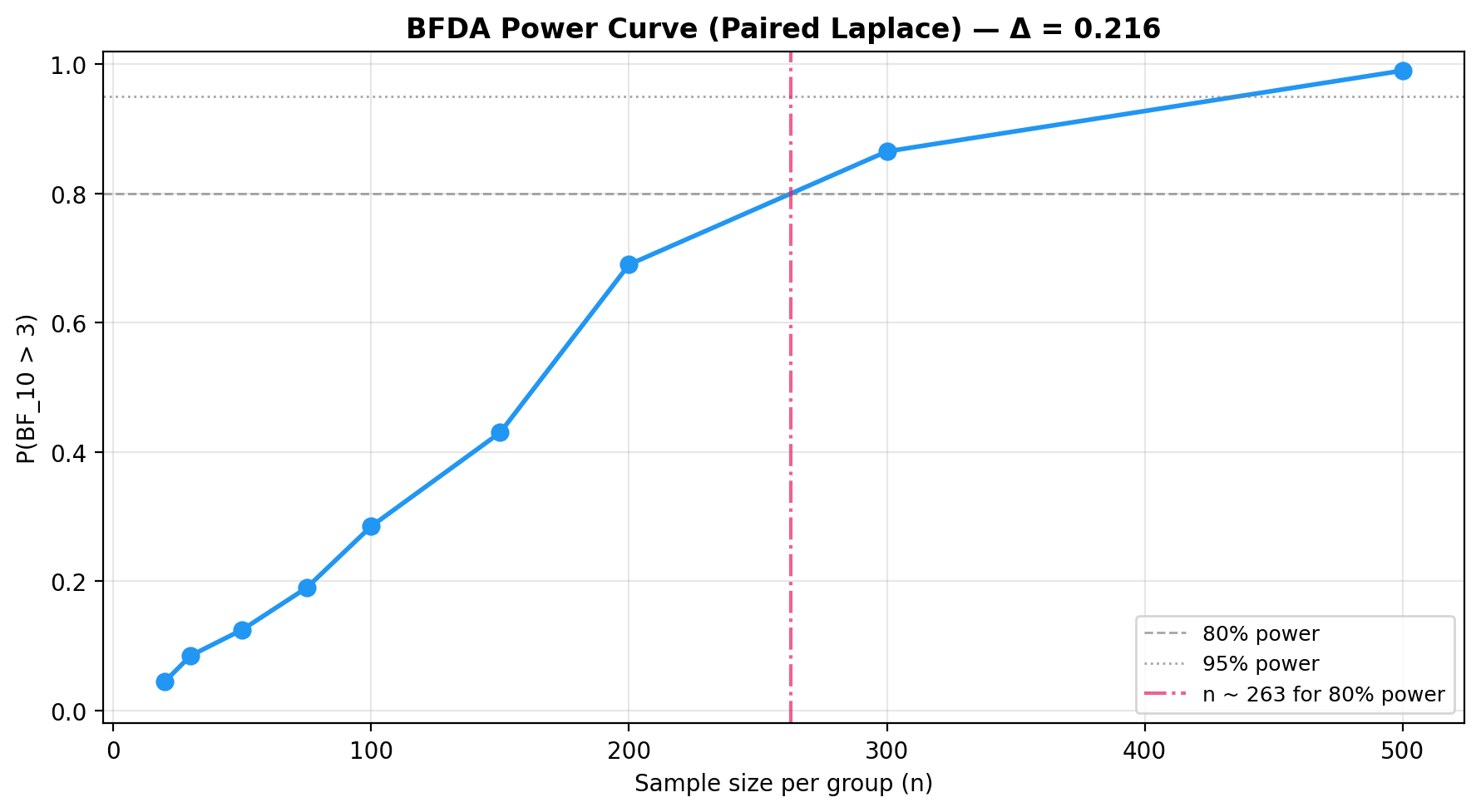
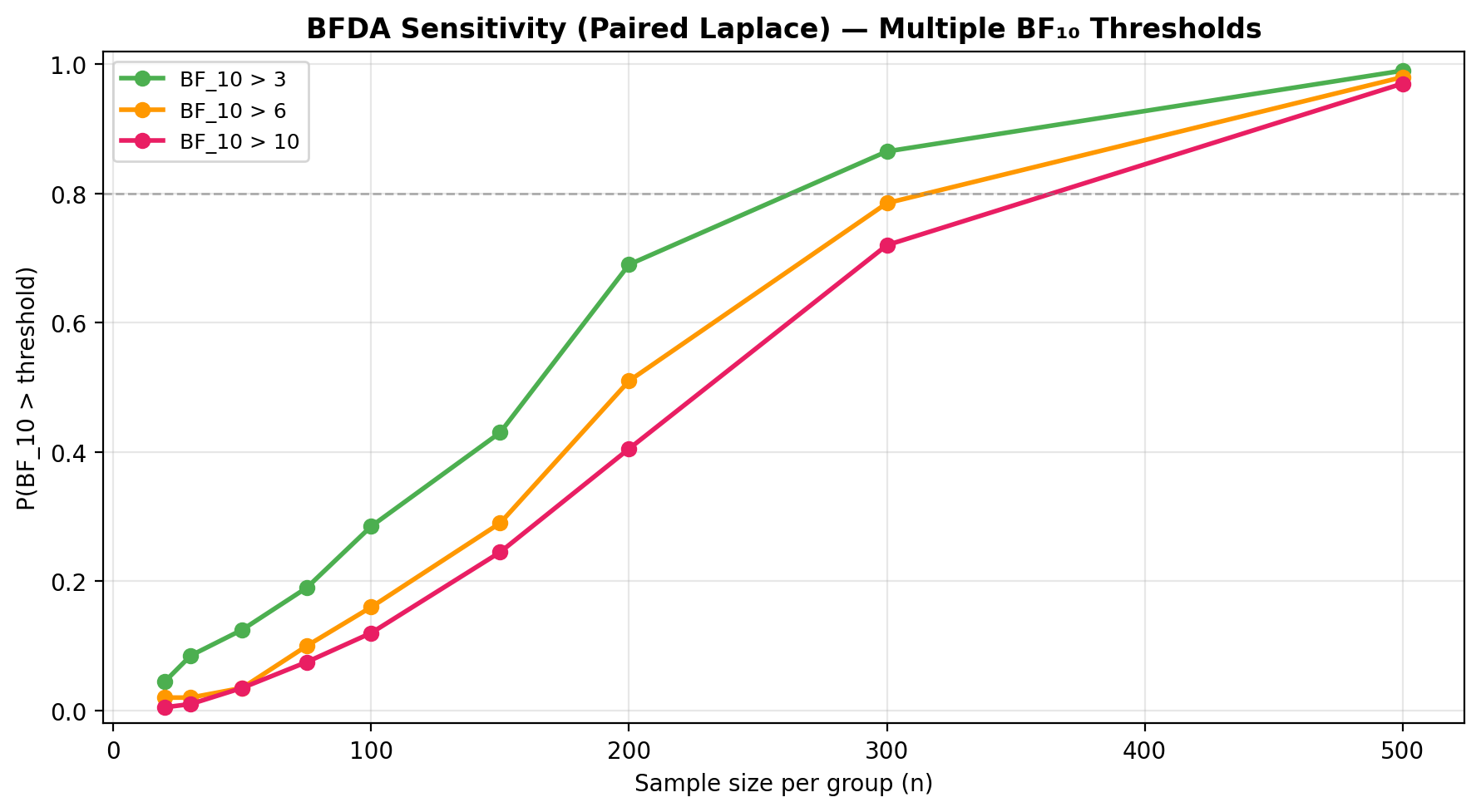
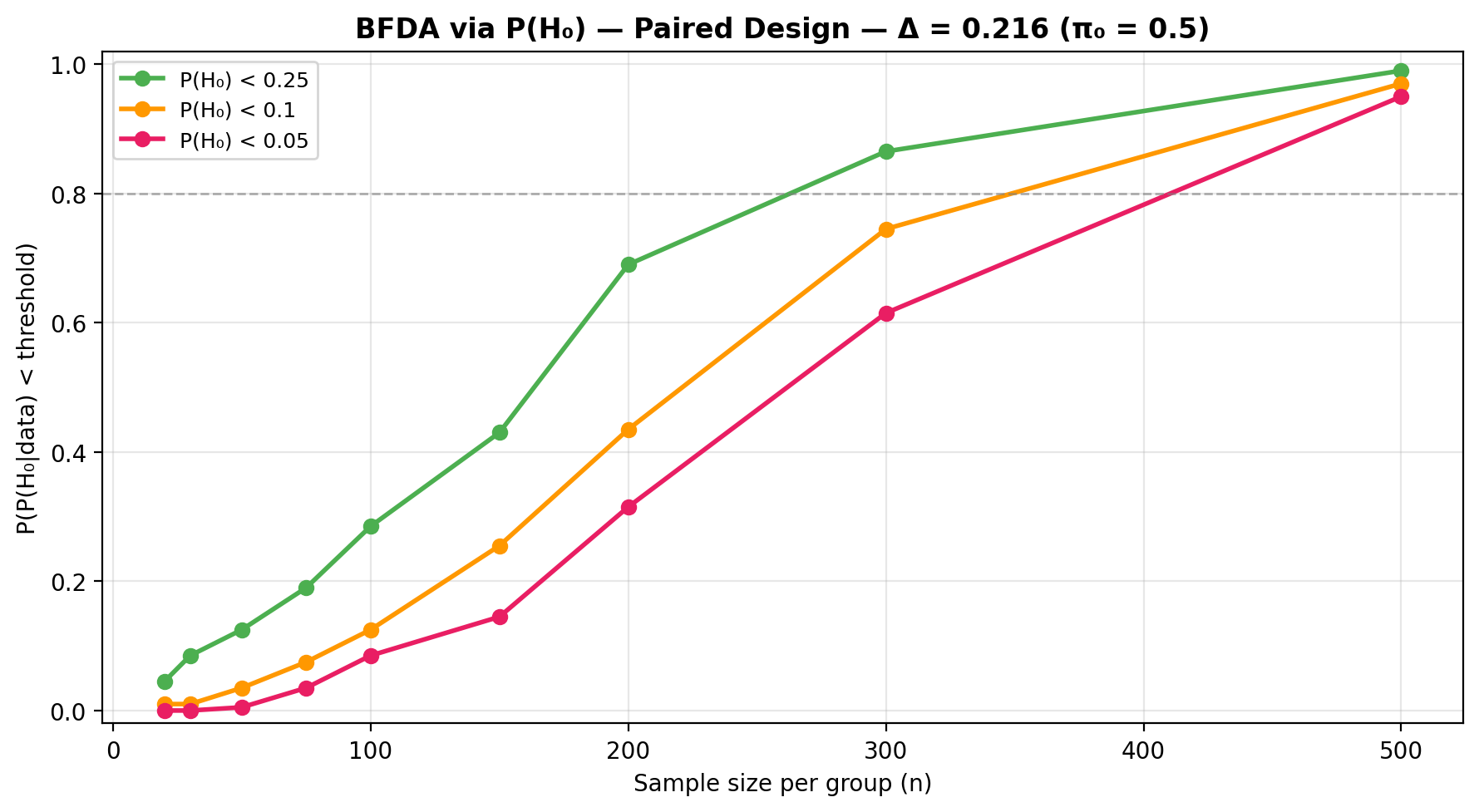
See the BFDA guide for sensitivity analysis and \(P(H_0)\)-based power curves.
Sequential design and decision making¶
In a sequential paired A/B test the binary observations arrive in batches over time and we update the Laplace posterior after each look. The pooled Bernoulli logistic likelihood depends on the data only through the four sufficient statistics \((n_A, k_A, n_B, k_B)\), so the cumulative counts carry all the information needed to recompute the Savage-Dickey Bayes factor on \(\delta_A = 0\), the posterior probability \(P(p_A > p_B)\) on the probability scale, and a ROPE decision at every look. Refitting on the running counts therefore yields exactly the same Laplace posterior as fitting all accumulated data in one shot — streaming introduces no additional approximation on top of the Laplace step itself.
Each refit is a damped Newton solve in 2D warm-started from the previous MAP, which typically converges in 1-3 iterations.
Stopping rule¶
At each look \(t\) the test evaluates the running \(\text{BF}_{10}^{(t)}\) and stops as soon as one of the following holds:
- \(\text{BF}_{10}^{(t)} \ge B_U\) (
bf_upper) -> stop for \(H_1\) (evidence of a difference). - \(\text{BF}_{10}^{(t)} \le B_L\) (
bf_lower) -> stop for \(H_0\) (evidence of practical equivalence). - \(\min(n_A^{(t)}, n_B^{(t)}) \ge n_{\max}\) -> stop because the budget is exhausted.
Because the Laplace posterior is a coherent likelihood-based object, optional stopping is permitted: performing many looks does not inflate a frequentist Type-I rate the way repeated \(p\)-values would.
Example: streaming paired Bernoulli batches¶
Ground truth on the logit scale: \(\mu = 0.5\), \(\delta_A = 0.6\). Each look delivers a batch of 25 paired observations.
import numpy as np
from bayesprop.resources.bayes_paired_laplace import SequentialPairedBayesPropTest
rng = np.random.default_rng(42)
p_A_true, p_B_true = 0.75, 0.62
def stream(n_batches: int = 40, batch_size: int = 25):
for _ in range(n_batches):
yield (
rng.binomial(1, p_A_true, size=batch_size),
rng.binomial(1, p_B_true, size=batch_size),
)
seq = SequentialPairedBayesPropTest(
prior_sigma_delta=1.0,
bf_upper=10.0,
bf_lower=0.1,
n_max=1000,
)
final = seq.run(stream())
print("Stopped:", seq.stopped, "after", len(seq.history), "looks")
print("Reason :", seq.stop_reason)
Inspect the final snapshot and history¶
The last SequentialLaplaceLookResult exposes the same diagnostics as
the batch test (Laplace posterior state, \(P(p_A > p_B)\), Savage-Dickey
BF, ROPE), and history_frame() returns one row per look:
ps = final.posterior_state
print(f"MAP: mu={ps.mu_map:.4f}, delta_A={ps.delta_A_map:.4f}")
print(f"P(p_A > p_B) = {final.P_A_greater_B:.4f}")
print(f"BF10 = {final.decision.bayes_factor.BF_10:.3g}")
print(f"ROPE decision: {final.decision.rope.decision}")
df = seq.history_frame() # per-look DataFrame
seq.plot_trajectory() # BF10 and P(A>B) vs cumulative n
Equivalence to a single-shot fit¶
Because the Laplace posterior depends only on the cumulative
sufficient statistics, fitting all accumulated data in one shot yields
the same MAP and covariance as the sequential refit at the final
look — i.e. seq.last_model matches a PairedBayesPropTest().fit(...)
on the materialised cumulative arrays.
See the runnable notebook at
src/notebooks/sequential_paired_laplace_demo.ipynb for the full demo.
API¶
See API Reference — Paired Model (Laplace) for full method documentation.